Science
Guided by data. Verified in the lab.
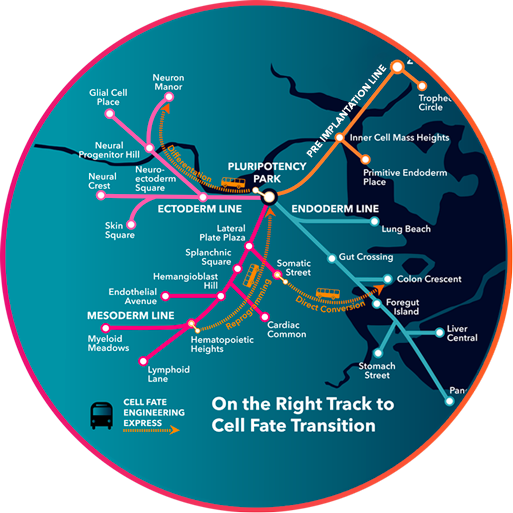
Our platform combines single-cell genomics, machine learning, and experimental biology to understand how cell identity is established and maintained, and then designs precise interventions to engineer it toward a defined destination.
We think of this as building a GPS for cell engineering, a system that maps where a cell is, identifies where it needs to go, and charts the most efficient route to get there.
Enabling Technologies for High Fidelity Cell Engineering
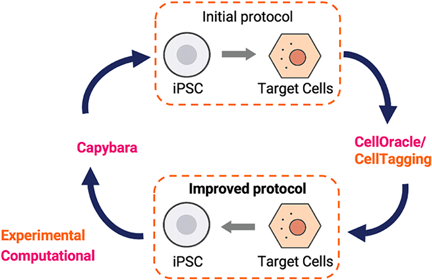
The CapyBio platform consists of four key building blocks: single-cell RNA sequencing maps the cell state landscape, CellOracle and CellTag chart the most efficient route towards the target state.
These predictions are tested experimentally in the lab, after which Capybara quantifies how closely the engineered cells match their primary human counterparts.
Together, they create a closed feedback system that iteratively improves the fidelity and scalability of engineered cells. For full references and peer-reviewed validation of our platform, visit our publications page.
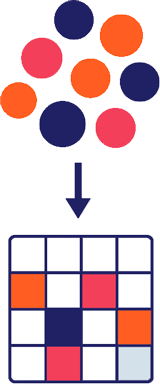
Single cell RNA sequencing
Every cell carries a unique molecular signature that encodes its identity. Traditional bulk assays average signals from mixed populations of cells and mask heterogeneity. Single-cell sequencing resolves this complexity by profiling thousands of individual cells in parallel. These data lie at the heart of our cell engineering platform. They let us to map distinct cell states both within primary human tissue and as cells are engineered toward a desired state. This knowledge becomes the blueprint for creating new cells and for evaluating whether they truly reflect their human counterparts.
CellOracle – in silico modeling
Current approaches to cell engineering rely heavily on trial-and-error. CellOracle changes that by modeling how cells respond to transcription factor perturbations in silico. It uses machine learning to build cell-state specific Gene Regulatory Networks (GRNs) and propagates gene perturbations through these learned networks to understand how they change cell identity, both at an individual cell and on a population level. This identifies factors which most efficiently drive cells toward a target identity, reducing wasted experiments and accelerating the discovery of effective interventions.
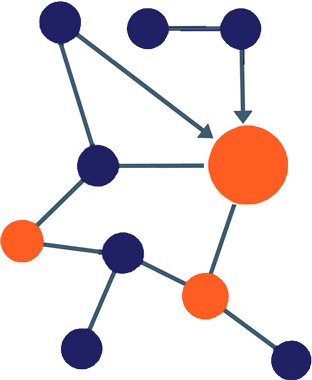
CellTagging – lineage tracing
In reprogramming and differentiation, cell fates are often specified before distinct cell states emerge. CellTagging is a single cell lineage tracing method that allows us to identify specific transcriptional and epigenetic features that uniquely mark cells that successfully reprogram to the desired state, thus allowing us to design experimental interventions to increase the yield of successfully reprogramming cells.
Capybara – Cell identity Scoring
Capybara uses quadratic programming to benchmark engineered cells against their primary human counterparts. By reporting compositional identity scores, it quantifies how faithfully each cell recapitulates the target cell type and detects the presence of any off-target identities. This quantitative, transcriptome-wide identity framework provides an objective, unbiased measure of success, enabling systematic optimization of cell engineering protocols.
Existing Approaches

- The differentiation process is rudimentary and unguided.
- Identity of final cells is difficult to measure.
- Few, if any, cells are converted to the target cell type.
The CapyBio Approach
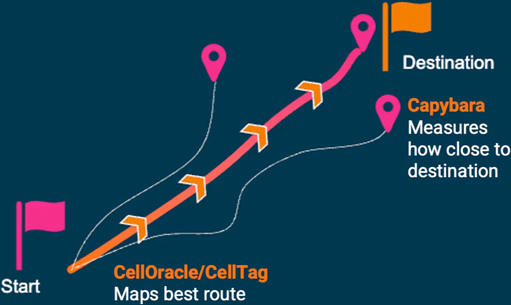
- CellOracle predicts factors for efficient conversion.
- Capybara quantifies cell identity, enabling iterative improvement.
- Greater purity of cell products at scale.
